Welcome !
Introduction
This R Markdown Website presents a single-cell RNA-sequencing experiment and analysis of dendritic cells following Salmonella infection.
Background
Salmonella Typhimurium (STM) is generally responsible for localized, self-limiting, gastroenteritis in humans. However, the multi-drug resistant S. Typhimurium ST313 pathovar has emerged across sub-Saharan Africa as a major cause of lethal bacteremia in children and HIV-infected adults. The isolate STM-D23580 is a representative blood-stream clinical isolate from a Malawian child, and demonstrates genome degradation resembling that of the human restricted pathogen S. Typhi. Dendritic cells (DCs) play an essential role in the initiation and establishment of antigen-specific immune responses. Modulation of DC functions by Salmonella has been reported as a mechanism to avoid adaptive immunity. Studies aimed at elucidating the interaction between invasive Salmonella and human DCs yielded important insights, yet they are limited by population-level measurements that mask fundamental differences among individual cells.
Method & Results
We combined single-cell RNA-sequencing technology with fluorescent labelling of bacteria to monitor gene expression variation among otherwise seemingly identical cells with regard to their infection condition. We quantified the early time course of gene expression induced by STM-LT2 or STM-D23580 infection in 373 human monocyte-derived dendritic cells. We demonstrated that transcriptional profiles could distinguish between different cellular infection phenotypes:
- One set of genes (involving the NF-kB signalling pathway with concomitant chemokine and cytokine production) responded simply to exposure to extracellular bacteria.
- Meanwhile, a different set of genes enriched for antigen presentation and proteolytic processes was markedly induced in Salmonella containing cells.
- Moreover, we found that gene expression heterogeneity between cells stimulated by extracellular or intracellular cues was increased during stimulation with invasive STM-D23580.
- Finally, we speculate that, in a subset of infected cells, STM-D23580 may impair the costimulatory signalling pathway by induction of IL10 and subversion of the CD83/MARCH1 axis.
Conclusion
Our data suggest that STM-D23580 strain may shape a different cellular environment in order to facilitate immune evasion. To our knowledge, this is the first single-cell RNA-sequencing study carried out in human DCs to provide new insights into the molecular contest at the Salmonella-host interface and suggest new areas of research to understand the mechanisms of invasive Salmonella infection.
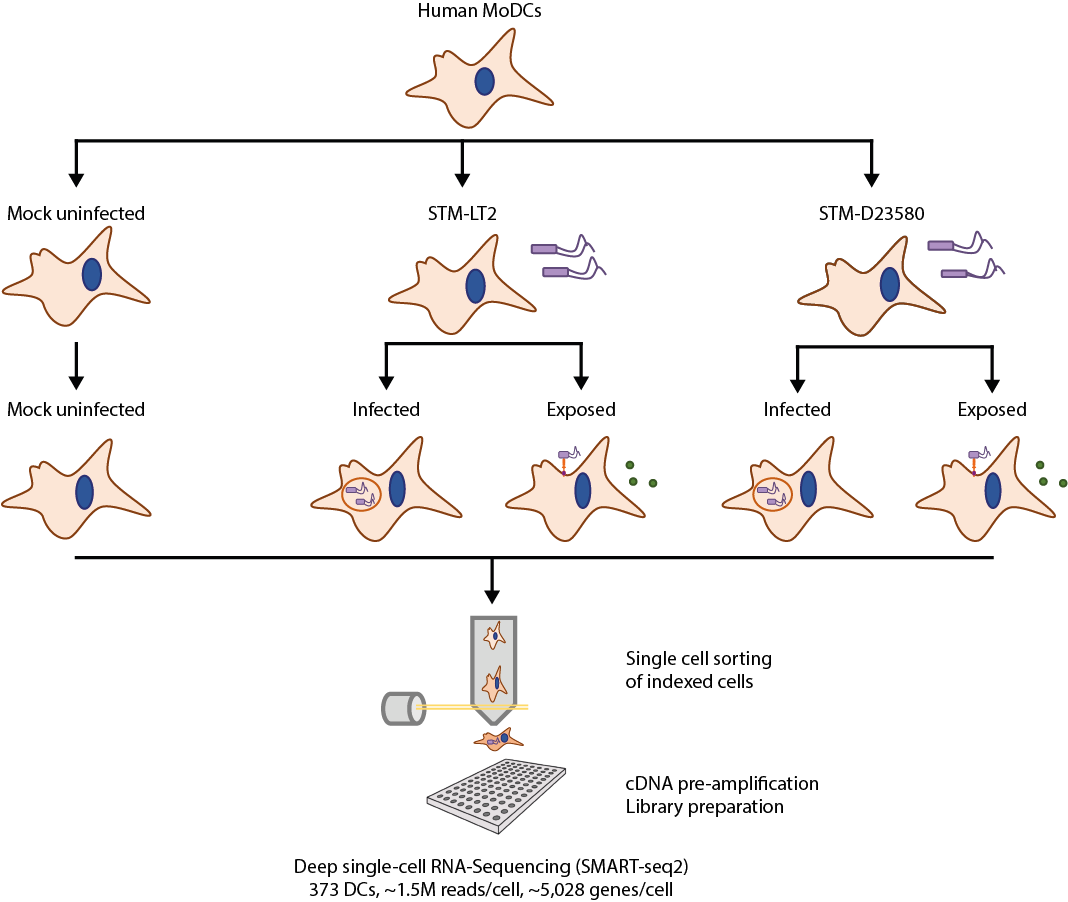
Experimental design